Antibiotic sensitivity testing
Antibiotic sensitivity testing or antibiotic susceptibility testing is the measurement of the susceptibility of bacteria to antibiotics. It is used because bacteria may have resistance to some antibiotics. Sensitivity testing results can allow a clinician to change the choice of antibiotics from empiric therapy, which is when an antibiotic is selected based on clinical suspicion about the site of an infection and common causative bacteria, to directed therapy, in which the choice of antibiotic is based on knowledge of the organism and its sensitivities.[1]
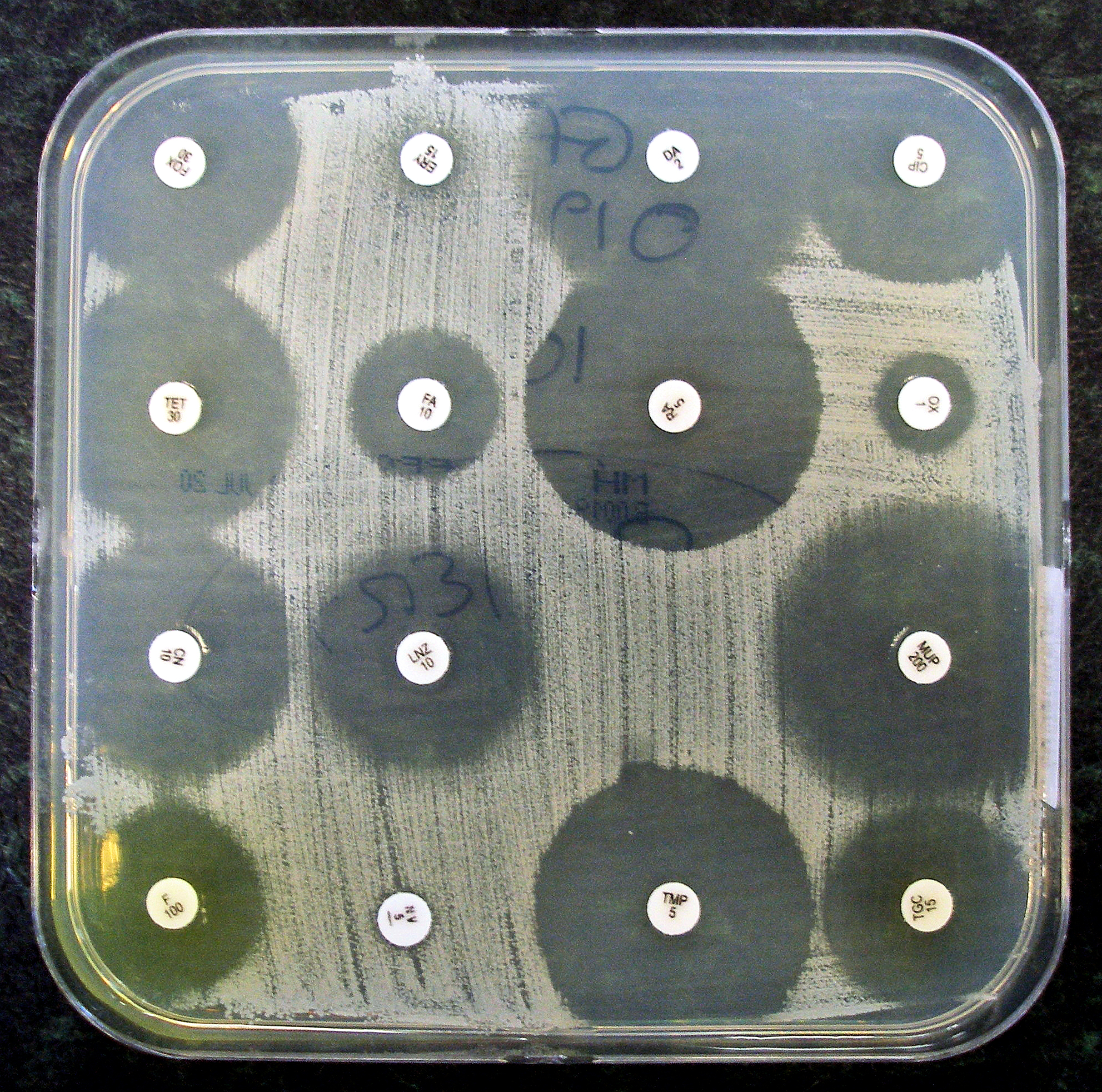
Sensitivity testing usually occurs in a medical laboratory, and uses culture methods that expose bacteria to antibiotics, or genetic methods that test to see if bacteria have genes that confer resistance. Culture methods often involve measuring the diameter of areas without bacterial growth, called zones of inhibition, around paper discs containing antibiotics on agar culture dishes that have been evenly inoculated with bacteria. The minimum inhibitory concentration, which is the lowest concentration of the antibiotic that stops the growth of bacteria, can be estimated from the size of the zone of inhibition.
Antibiotic susceptibility testing has been needed since the discovery of the beta-lactam antibiotic penicillin. Initial methods were phenotypic, and involved culture or dilution. The Etest, an antibiotic impregnated strip, has been available since the 1980s, and genetic methods such as polymerase chain reaction (PCR) testing have been available since the early 2000s. Research is ongoing into improving current methods by making them faster or more accurate, as well as developing new methods for testing, such as microfluidics.
Uses[edit]
In clinical medicine, antibiotics are most frequently prescribed on the basis of a person's symptoms and medical guidelines. This method of antibiotic selection is called empiric therapy,[1] and it is based on knowledge about what bacteria cause an infection, and to what antibiotics bacteria may be sensitive or resistant.[1] For example, a simple urinary tract infection might be treated with trimethoprim/sulfamethoxazole.[2] This is because Escherichia coli is the most likely causative bacterium, and may be sensitive to that combination antibiotic.[2] However, bacteria can be resistant to several classes of antibiotics.[2] This resistance might be because a type of bacteria has intrinsic resistance to some antibiotics,[2] because of resistance following past exposure to antibiotics,[2] or because resistance may be transmitted from other sources such as plasmids.[3] Antibiotic sensitivity testing provides information about which antibiotics are more likely to be successful and should therefore be used to treat the infection.[1]
Antibiotic sensitivity testing is also conducted at a population level in some countries as a form of screening.[4] This is to assess the background rates of resistance to antibiotics (for example with methicillin-resistant Staphylococcus aureus), and may influence guidelines and public health measures.[4]
Reporting[edit]
Bacteria are marked as sensitive, resistant, or having intermediate resistance to an antibiotic based on the minimum inhibitory concentration (MIC), which is the lowest concentration of the antibiotic that stops the growth of bacteria. The MIC is compared to standard threshold values (called "breakpoints") for a given bacterium and antibiotic.[27] Breakpoints for the same organism and antibiotic may differ based on the site of infection:[28] for example, the CLSI generally defines Streptococcus pneumoniae as sensitive to intravenous penicillin if MICs are ≤0.06 μg/ml, intermediate if MICs are 0.12 to 1 μg/ml, and resistant if MICs are ≥2 μg/ml, but for cases of meningitis, the breakpoints are considerably lower.[29] Sometimes, whether an antibiotic is marked as resistant is also based on bacterial characteristics that are associated with known methods of resistance such as the potential for beta-lactamase production.[27][20] Specific patterns of drug resistance or multidrug resistance may be noted, such as the presence of an extended-spectrum beta lactamase.[27] Such information may be useful to the clinician, who can change the empiric treatment to a tailored treatment that is directed only at the causative bacterium.[1][9] The results of antimicrobial susceptibility tests performed during a given time period can be compiled, usually in the form of a table, to form an antibiogram.[30][31] Antibiograms help the clinician to select the best empiric antimicrobial therapy based on the local resistance patterns until the laboratory test results are available.[31]
History[edit]
Since the discovery of the beta-lactam antibiotic penicillin, the rates of antimicrobial resistance have increased.[37] Over time, methods for testing the sensitivity of bacteria to antibiotics have developed and changed.[25]
Alexander Fleming in the 1920s developed the first method of susceptibility testing. The "gutter method" that he developed was a diffusion method, involving an antibiotic that was diffused through a gutter made of agar.[25] In the 1940s, multiple investigators, including Pope, Foster and Woodruff, Vincent and Vincent used paper discs instead.[25] All these methods involve testing only susceptibility to penicillin.[25] The results were difficult to interpret and not reliable, because of inaccurate results that were not standardised between laboratories.[25]
Dilution has been used as a method to grow and identify bacteria since the 1870s, and as a method of testing the susceptibility of bacteria to antibiotics since 1929, also by Alexander Fleming.[25] The way of determining susceptibility changed from how turbid the solution was, to the pH (in 1942), to optical instruments.[25] The use of larger tube-based "macrodilution" testing has been superseded by smaller "microdilution" kits.[5]
In 1966, the World Health Organisation confirmed the Kirby-Bauer method as the standard method for susceptibility testing; it is simple, cost-effective and can test multiple antibiotics.[25]
The Etest was developed in 1980 by Bolmstrӧm and Eriksson, and MALDI-TOF developed in 2000s.[25] An array of automated systems has been developed since and after the 1980s.[25] PCR was the first genetic test available and first published as a method of detecting antibiotic susceptibility in 2001.[25]
Further research[edit]
Point-of-care testing is being developed to speed up the time for testing, and to help practitioners avoid prescribing unnecessary antibiotics in the style of precision medicine.[38] Traditional techniques typically take between 12 and 48 hours,[6] although it can take up to five days.[27] In contrast, rapid testing using molecular diagnostics is defined as "being feasible within an 8-h(our) working shift".[6] Progress has been slow due to a range of reasons including cost and regulation.[39]
Additional research is focused at the shortcomings of current testing methods. As well as the duration it takes to report phenotypic methods, they are laborious, have difficult portability and are difficult to use in resource-limited settings, and have a chance of cross-contamination.[25]
As of 2017, point-of-care resistance diagnostics were available for methicillin-resistant Staphylococcus aureus (MRSA), rifampin-resistant Mycobacterium tuberculosis (TB), and vancomycin-resistant enterococci (VRE) through GeneXpert by molecular diagnostics company Cepheid.[40]
Quantitative PCR, with the view of determining the percent of a detected bacteria that possesses a resistance gene, is being explored.[9] Whole genome sequencing of isolated bacteria is also being explored, and likely to become more available as costs decrease and speed increases over time.[9]
Additional methods explored include microfluidics, which uses a small amount of fluid and a variety of testing methods, such as optical, electrochemical, and magnetic.[9] Such assays do not require much fluid to be tested, are rapid and portable.[9]
The use of fluorescent dyes has been explored.[9] These involve labelled proteins targeted at biomarkers, nucleic acid sequences present within cells that are found when the bacterium is resistant to an antibiotic.[9] An isolate of bacteria is fixed in position and then dissolved. The isolate is then exposed to fluorescent dye, which will be luminescent when viewed.[9]
Improvements to existing platforms are also being explored, including improvements in imaging systems that are able to more rapidly identify the MIC in phenotypic samples; or the use of bioluminescent enzymes that reveal bacterial growth to make changes more easily visible.[25]


